Layman’s Introduction to KNN

k-nearest neighbour algorithm is where most people begin when starting with machine learning.
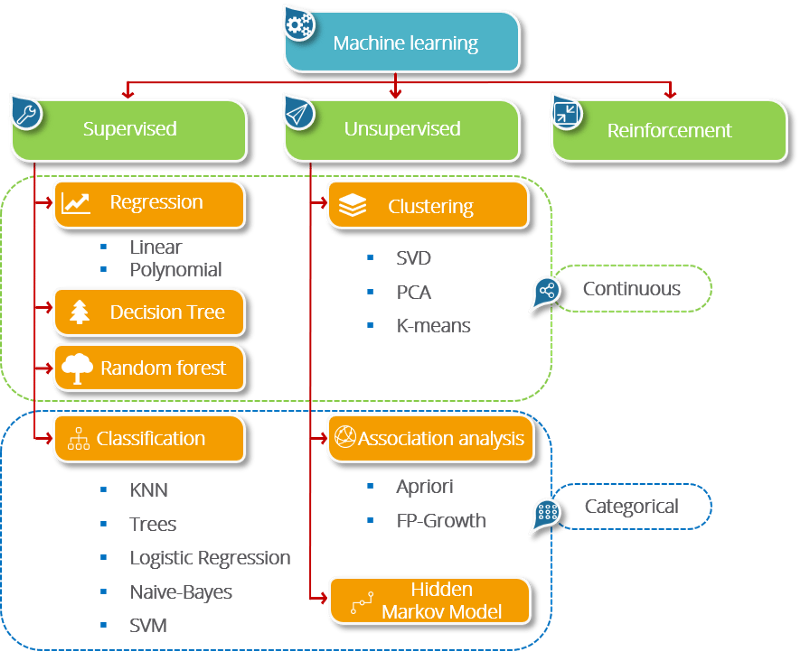
kNN stands for k-Nearest Neighbours. It is a supervised learning algorithm. This means that we train it under supervision. We train it using the labelled data already available to us. Given a labelled dataset consisting of observations (x,y), we would like to capture the relationship between x — the data and y — the label. More formally, we want to learn a function g : X→Y so that given an unseen observation X, g(x) can confidently predict the corresponding output Y.
Other examples of supervised learning algorithms include random forests, linear regression and logistic regression.
kNN is very simple to implement and is most widely used as a first step in any machine learning setup. It is often used as a benchmark for more complex classifiers such as Artificial Neural Networks (ANN) and Support Vector Machines (SVM). Despite its simplicity, k-NN can outperform more powerful classifiers and is used in a variety of applications such as economic forecasting, data compression and genetics.
- Genetics

- Agriculture
- Aviation — Air traffic flow prediction
As with most technological progress in the early 1900s, KNN algorithm was also born out of research done for the armed forces. Two offices of USAF School of Aviation Medicine — Fix and Hodges (1951) wrote a technical report introducing a non-parametric method for pattern classification that has since become popular as the k-nearest neighbor (kNN) algorithm.

How does it work?
Let’s say we have a dataset with two kinds of points — Label 1 and Label 2. Now given a new point in this dataset we want to figure out its label. The way it is done in kNN is by taking a majority vote of its k nearest neighbours. k can take any value between 1 and infinity but in most practical cases k is less than 30.
Blue Circles v/s Orange Triangles
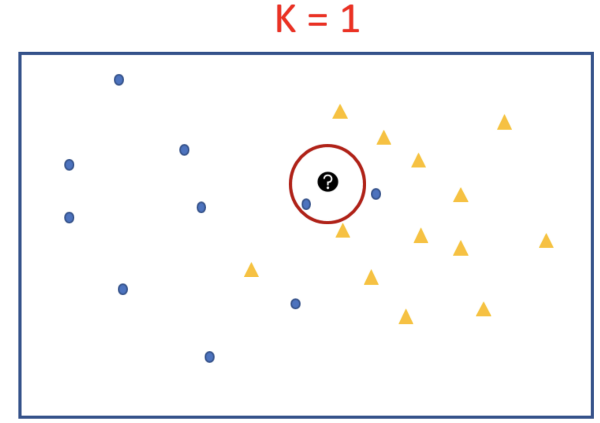
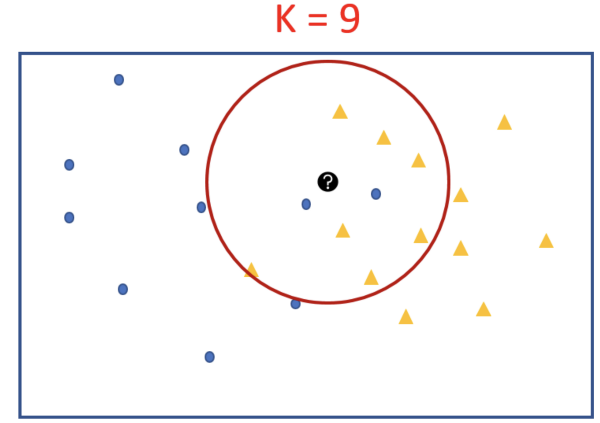
Let’s say we have two groups of points — blue-circles and orange-triangles. We want to classify the Test Point = black circle with a question mark, as either a blue circle or an orange triangle.
Goal: To label the black circle.
For K = 1 we will look at the first nearest neighbour. Since we take majority vote and there is only 1 voter we assign its label to our black test point. We can see that the test point will be classified as a blue circle for k=1.
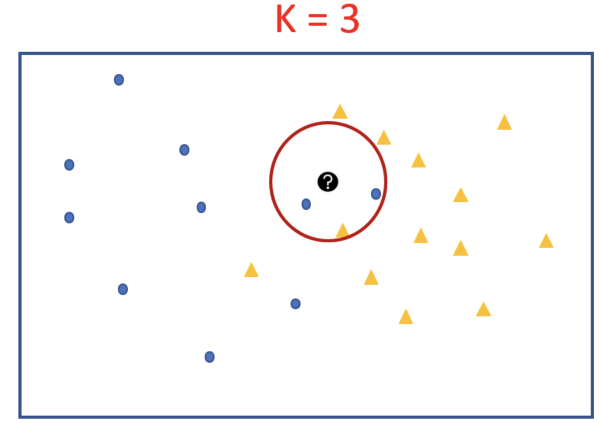
Expanding our search radius to K=3 also keeps the result same, except that this time it is not an absolute majority, it’s 2 out of 3. Still with k=3 test point is predicted to have the class blue-circle →because the majority of points are blue.
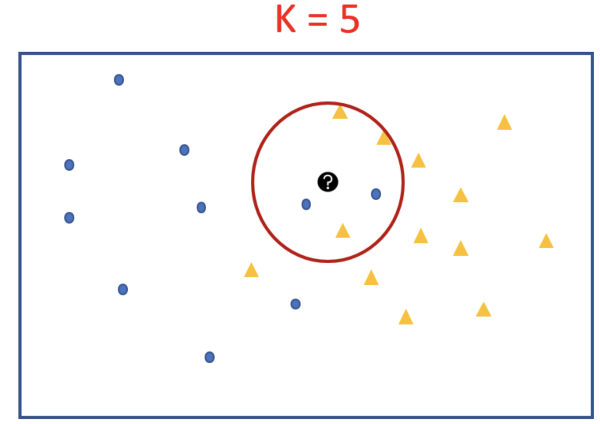
Let’s see how k=5 and K =9 do. To look at the nearest neighbors we draw circle with test point at the centre and stop when 5 points fall inside the circle.
When we look at the 5 and subsequently at K = 9, the majority of the closest neighbors of our test point are orange-triangles. That indicates that the test point must be an orange triangle.
Now that we have labelled this one test point we repeat the same process over all the unknown points (i.e. the test set). Once all test points are labelled using k-NN we try separating them using a decision boundary. Decision boundary shows how well the training set is separated.

That’s the gist of how k-NN happens. Let’s see it from the point of view of a machine learning engineer’s brain.
First they would choose k. We already saw above that a bigger k takes a vote of larger number of points. This means higher chances of being correct. But at what cost, you say?
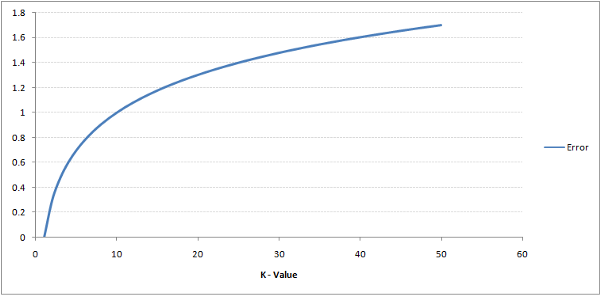
Getting the k nearest neighbours means sorting through the distances. That is a costly operation. A very high processing power is needed which translates to either longer processing time or costlier processor. Higher the K costlier the whole procedure. But too low a k would result in overfitting.
A very low k will fail to generalize. A very high k is costly.
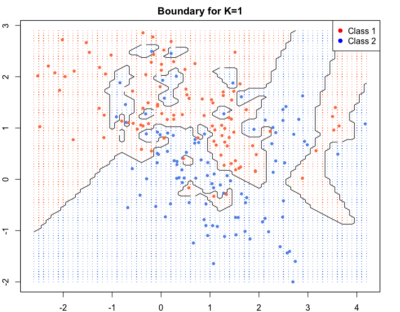
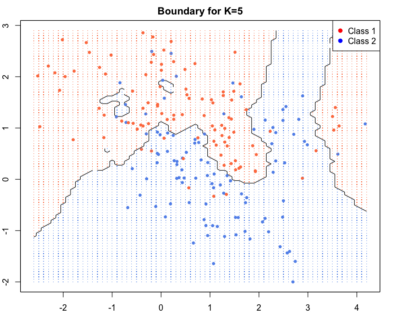
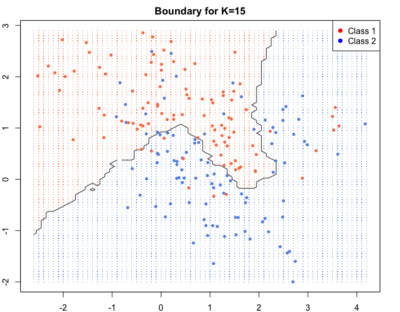
As we go to higher K’s the boundaries become smooth.Blue and red regions are broadly separated. Some blue and red soldiers are left behind the enemy lines. They are collateral damage. They account for loss in training accuracy but lead to better generalisation and high test accuracy i.e. high accuracy of correct labelling for new points.
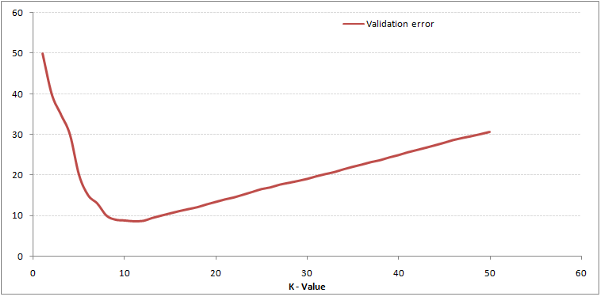
A graph of validation error when plotted against K would typically look like this. We can see that around K = 8 the error is minimum. It goes up on either side.
Then they would try to find distances between points. How do we decide which neighbours are near and which are not?
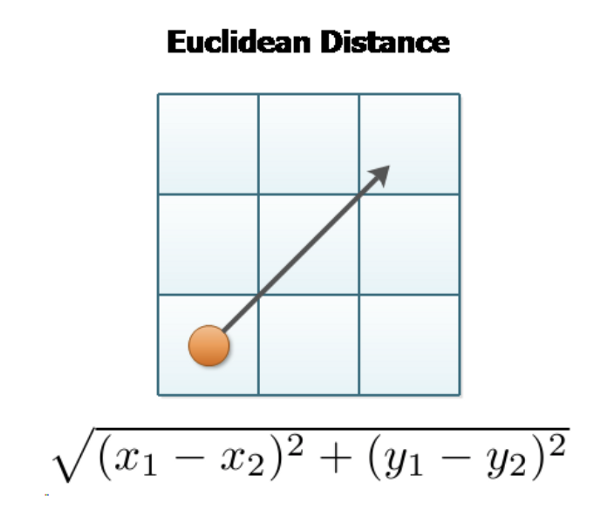
- Euclidean Distance — Most common distance metric
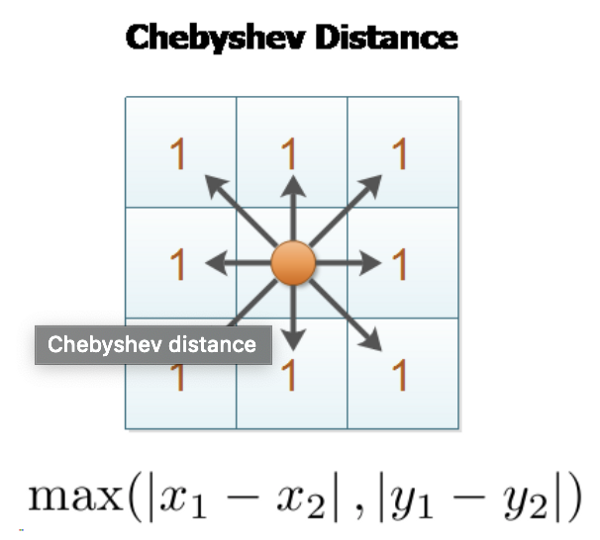
- Chebyshev Distance — L∞ Distance
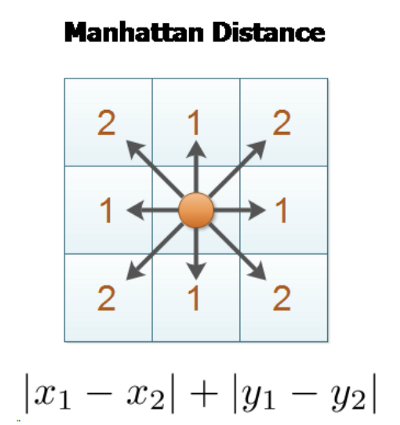
- Manhattan Distance — L1 Distance : Sum of the (absolute) differences of their coordinates.
In chess, the distance between squares on the chessboard for rooks is measured in Manhattan distance; kings and queens use Chebyshev distance — Wikipedia
Once we know how to compare points based on distance we would like to train our model. The best part about k-NN is that there is no explicit training step for it. We already know all that is to know about our dataset — its labels. In essence the training phase of the algorithm consists only of storing the feature vectors and class labels of the training samples.

K-NN is a lazy learner because it doesn’t learn a discriminative function from the training data but memorizes the training dataset instead.
An eager learner has a model fitting or training step. A lazy learner does not have a training phase.
Finally, why k-NN?
- Quick to implement : Which is why it is popular as a benchmarking algorithm.
- Less training time: Faster turn around time
- Comparable accuracies: Its prediction accuracy as indicated in a lot of research papers is fairly high for a lot of applications.
k-NN is a life saver when one has to quickly deliver a solution with fairly accurate results. In most tools like MATLAB, python, R it is given as a single line command. Despite that it is very easy to implement and fun to try.
Source code for the R Plots
library(ElemStatLearn)
require(class)
#Data Extraction
KVAL <- 15
x <- mixture.example$x
g <- mixture.example$y
xnew <- mixture.example$xnew
#KNN and boundary extraction
mod15 <- knn(x, xnew, g, k=KVAL, prob=TRUE)
prob <- attr(mod15, "prob")
prob <- ifelse(mod15=="1", prob, 1-prob)
px1 <- mixture.example$px1
px2 <- mixture.example$px2
prob15 <- matrix(mod15, length(px1), length(px2))
#Plotting Boundary
par(mar=rep(2,4))
contour(px1, px2, prob15, levels=0.5, labels="", xlab="x", ylab="y", main="", axes=TRUE)
#Plotting red and blue points
points(x, pch = 20, col=ifelse(g==1, "coral", "cornflowerblue"))gd <- expand.grid(x=px1, y=px2)
points(gd, pch=".", cex=0.001, col=ifelse(prob15>0.5, "coral", "cornflowerblue"))
legend("topright", pch = c(19, 19), col = c( "red","blue"), legend = c( "Class 1", "Class 2"))
title(main="Boundary for K=15", sub="ii", xlab="Feature 1", ylab="Feature 2")
box()
Member discussion